Chapter 12 Bach mouse mammary gland (10X Genomics)
12.1 Introduction
This performs an analysis of the Bach et al. (2017) 10X Genomics dataset, from which we will consider a single sample of epithelial cells from the mouse mammary gland during gestation.
12.3 Quality control
is.mito <- rowData(sce.mam)$SEQNAME == "MT"
stats <- perCellQCMetrics(sce.mam, subsets=list(Mito=which(is.mito)))
qc <- quickPerCellQC(stats, percent_subsets="subsets_Mito_percent")
sce.mam <- sce.mam[,!qc$discard]colData(unfiltered) <- cbind(colData(unfiltered), stats)
unfiltered$discard <- qc$discard
gridExtra::grid.arrange(
plotColData(unfiltered, y="sum", colour_by="discard") +
scale_y_log10() + ggtitle("Total count"),
plotColData(unfiltered, y="detected", colour_by="discard") +
scale_y_log10() + ggtitle("Detected features"),
plotColData(unfiltered, y="subsets_Mito_percent",
colour_by="discard") + ggtitle("Mito percent"),
ncol=2
)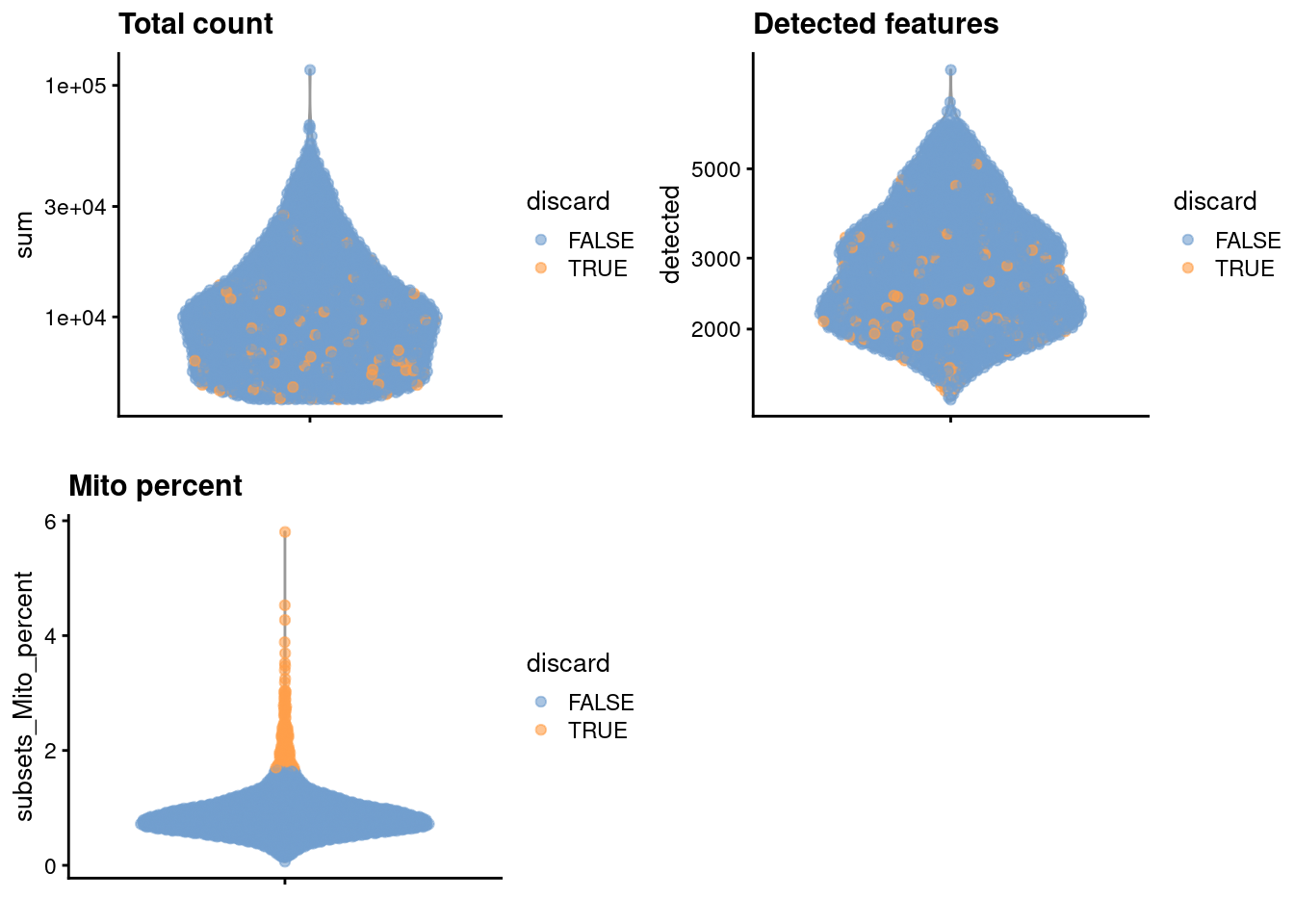
Figure 12.1: Distribution of each QC metric across cells in the Bach mammary gland dataset. Each point represents a cell and is colored according to whether that cell was discarded.
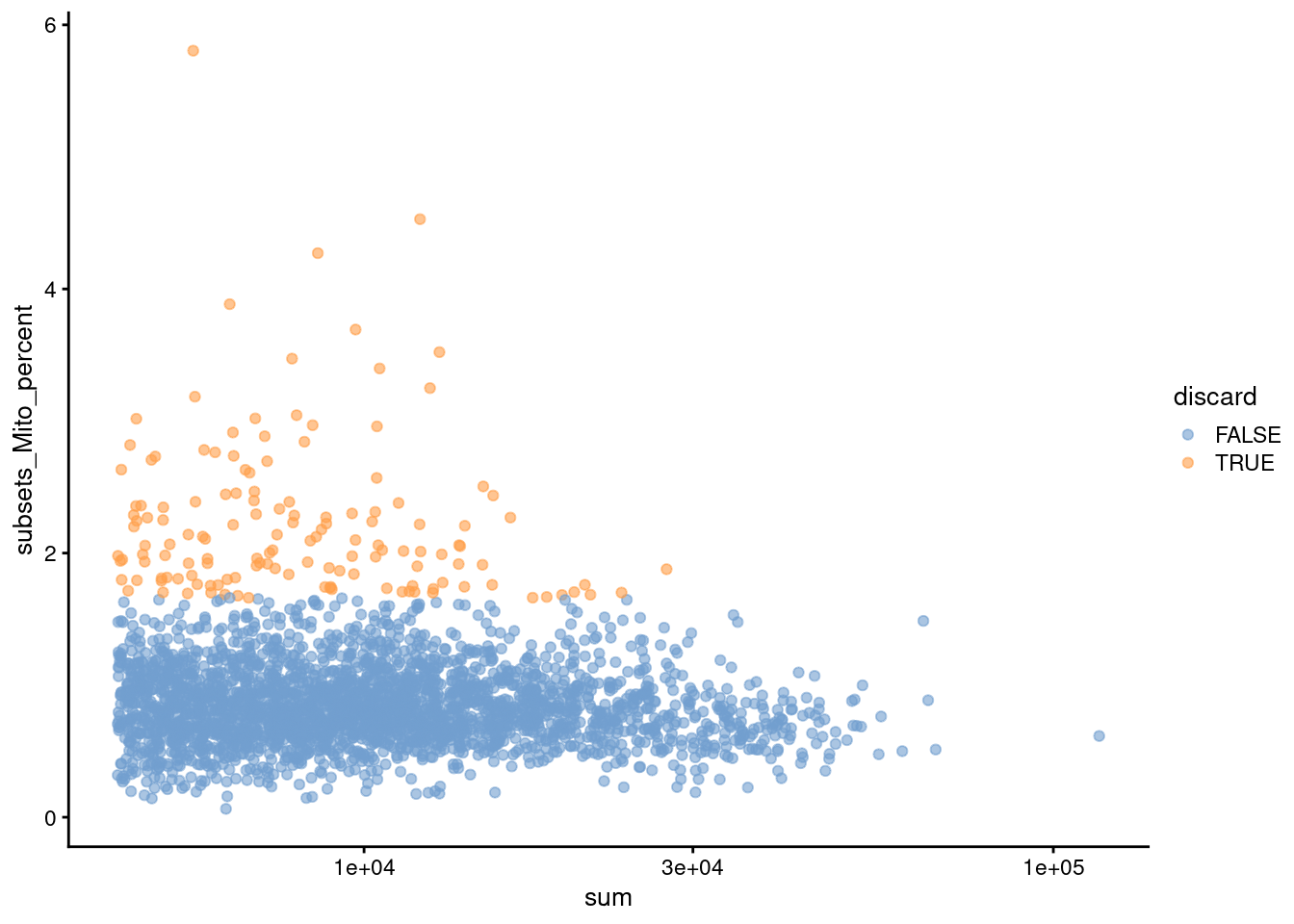
Figure 12.2: Percentage of mitochondrial reads in each cell in the Bach mammary gland dataset compared to its total count. Each point represents a cell and is colored according to whether that cell was discarded.
## low_lib_size low_n_features high_subsets_Mito_percent
## 0 0 143
## discard
## 14312.4 Normalization
library(scran)
set.seed(101000110)
clusters <- quickCluster(sce.mam)
sce.mam <- computeSumFactors(sce.mam, clusters=clusters)
sce.mam <- logNormCounts(sce.mam)## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 0.271 0.522 0.758 1.000 1.204 10.958plot(librarySizeFactors(sce.mam), sizeFactors(sce.mam), pch=16,
xlab="Library size factors", ylab="Deconvolution factors", log="xy")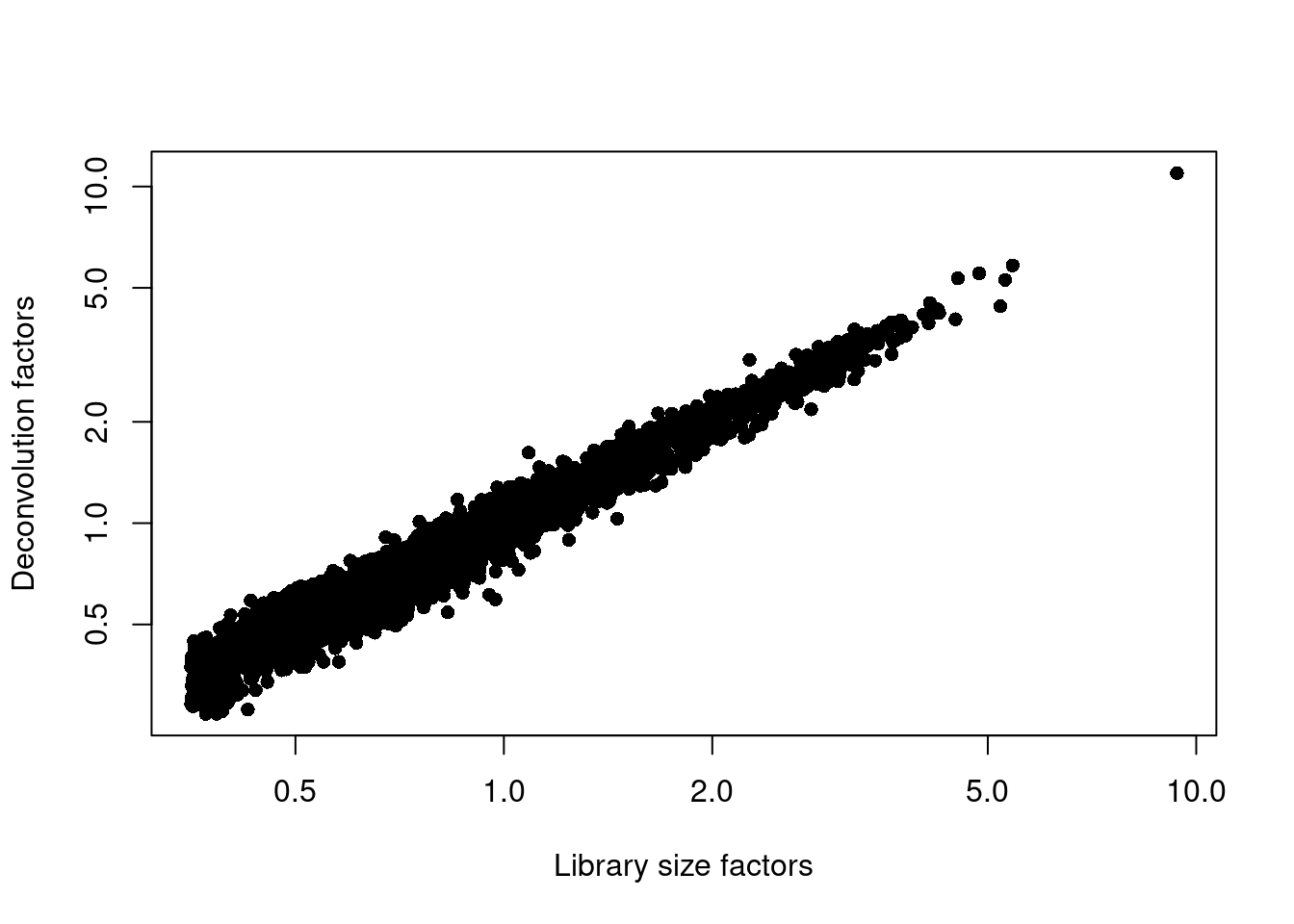
Figure 12.3: Relationship between the library size factors and the deconvolution size factors in the Bach mammary gland dataset.
12.5 Variance modelling
We use a Poisson-based technical trend to capture more genuine biological variation in the biological component.
set.seed(00010101)
dec.mam <- modelGeneVarByPoisson(sce.mam)
top.mam <- getTopHVGs(dec.mam, prop=0.1)plot(dec.mam$mean, dec.mam$total, pch=16, cex=0.5,
xlab="Mean of log-expression", ylab="Variance of log-expression")
curfit <- metadata(dec.mam)
curve(curfit$trend(x), col='dodgerblue', add=TRUE, lwd=2)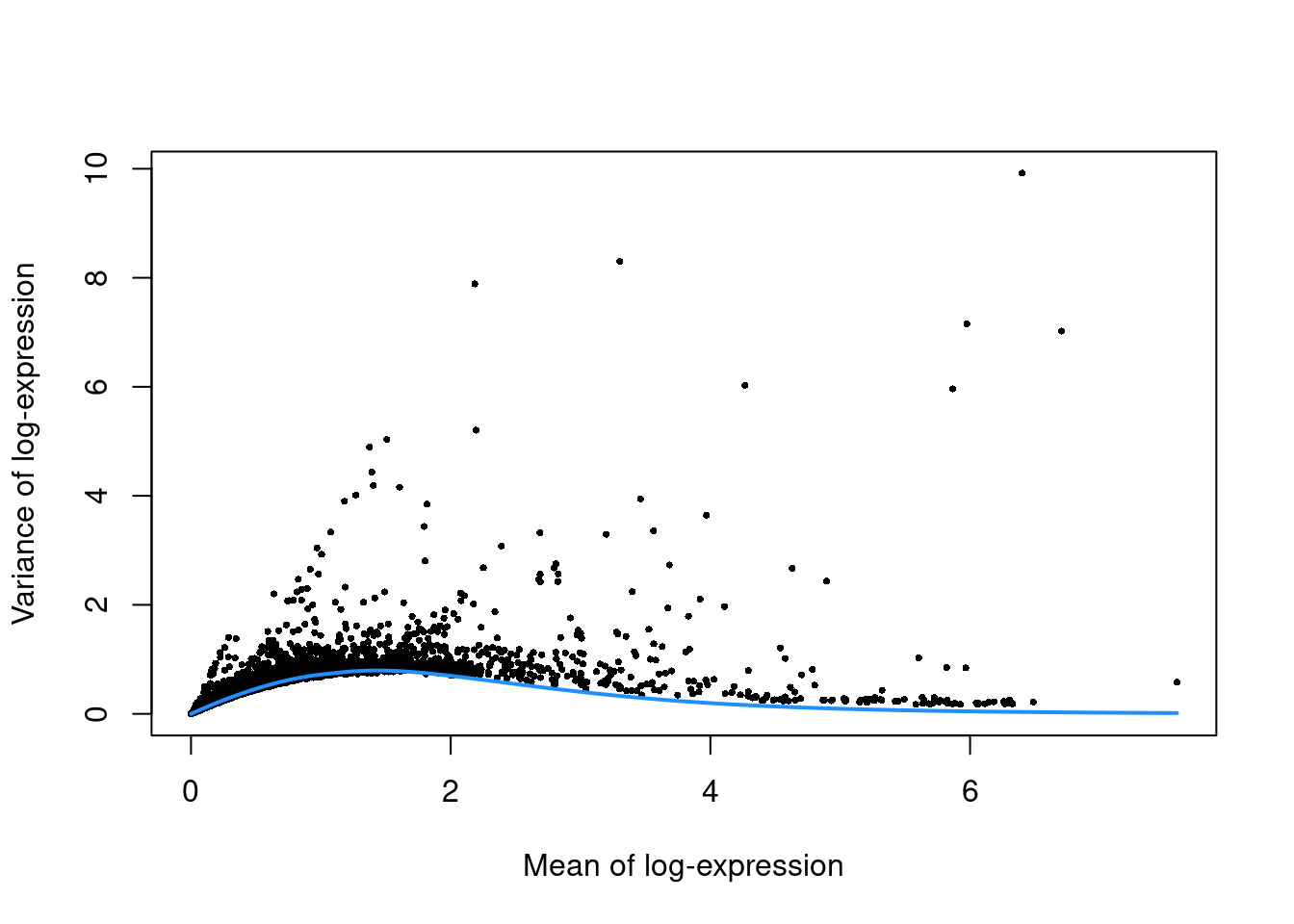
Figure 12.4: Per-gene variance as a function of the mean for the log-expression values in the Bach mammary gland dataset. Each point represents a gene (black) with the mean-variance trend (blue) fitted to simulated Poisson counts.
12.6 Dimensionality reduction
library(BiocSingular)
set.seed(101010011)
sce.mam <- denoisePCA(sce.mam, technical=dec.mam, subset.row=top.mam)
sce.mam <- runTSNE(sce.mam, dimred="PCA")## [1] 1512.7 Clustering
We use a higher k to obtain coarser clusters (for use in doubletCluster() later).
snn.gr <- buildSNNGraph(sce.mam, use.dimred="PCA", k=25)
colLabels(sce.mam) <- factor(igraph::cluster_walktrap(snn.gr)$membership)##
## 1 2 3 4 5 6 7 8 9 10
## 550 799 716 452 24 84 52 39 32 24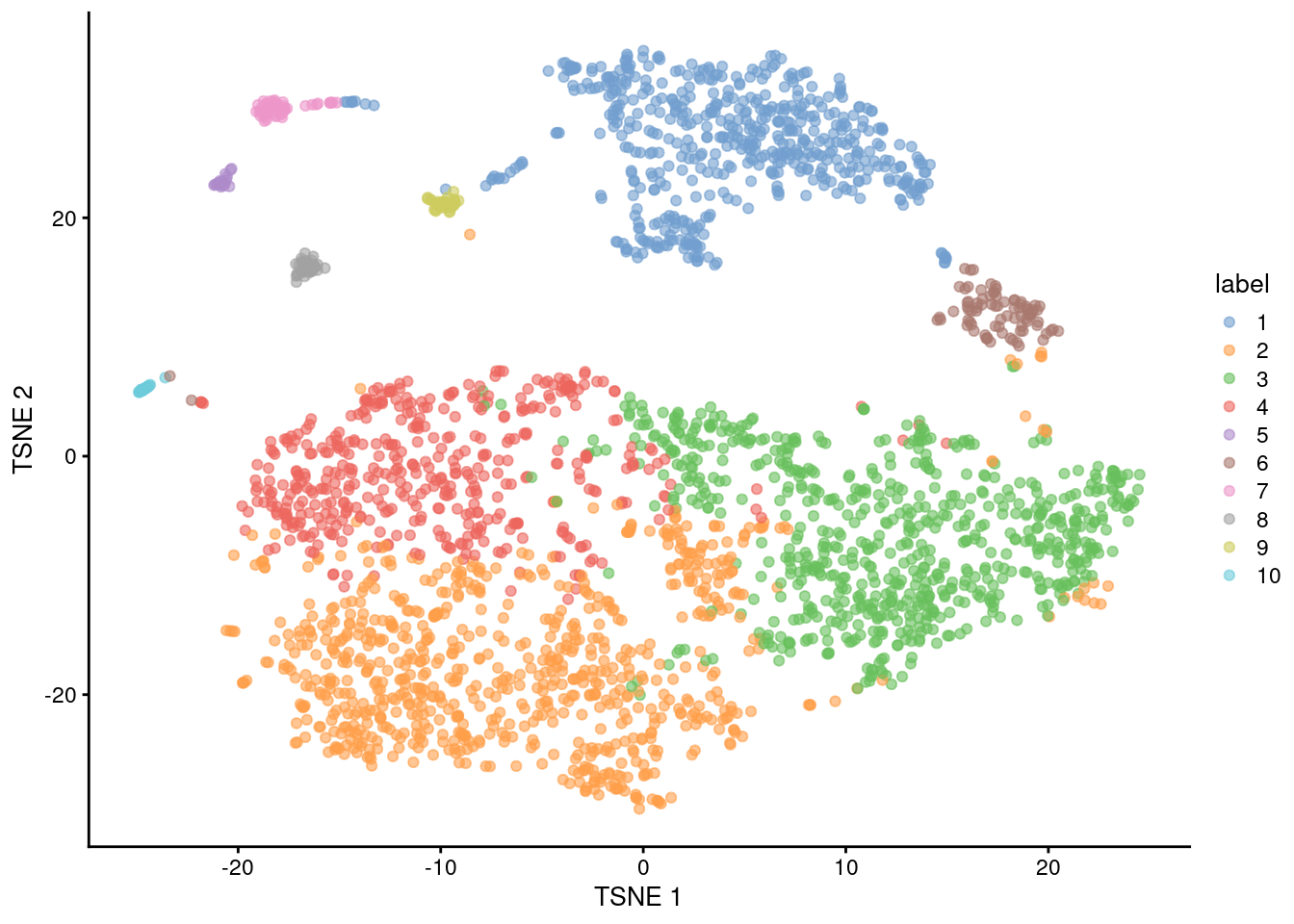
Figure 12.5: Obligatory \(t\)-SNE plot of the Bach mammary gland dataset, where each point represents a cell and is colored according to the assigned cluster.
Session Info
R version 4.3.1 (2023-06-16)
Platform: x86_64-pc-linux-gnu (64-bit)
Running under: Ubuntu 22.04.3 LTS
Matrix products: default
BLAS: /home/biocbuild/bbs-3.18-bioc/R/lib/libRblas.so
LAPACK: /usr/lib/x86_64-linux-gnu/lapack/liblapack.so.3.10.0
locale:
[1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_GB LC_COLLATE=C
[5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
[7] LC_PAPER=en_US.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
time zone: America/New_York
tzcode source: system (glibc)
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] BiocSingular_1.18.0 scran_1.30.0
[3] AnnotationHub_3.10.0 BiocFileCache_2.10.0
[5] dbplyr_2.3.4 scater_1.30.0
[7] ggplot2_3.4.4 scuttle_1.12.0
[9] ensembldb_2.26.0 AnnotationFilter_1.26.0
[11] GenomicFeatures_1.54.0 AnnotationDbi_1.64.0
[13] Matrix_1.6-1.1 scRNAseq_2.15.0
[15] SingleCellExperiment_1.24.0 SummarizedExperiment_1.32.0
[17] Biobase_2.62.0 GenomicRanges_1.54.0
[19] GenomeInfoDb_1.38.0 IRanges_2.36.0
[21] S4Vectors_0.40.0 BiocGenerics_0.48.0
[23] MatrixGenerics_1.14.0 matrixStats_1.0.0
[25] BiocStyle_2.30.0 rebook_1.12.0
loaded via a namespace (and not attached):
[1] rstudioapi_0.15.0 jsonlite_1.8.7
[3] CodeDepends_0.6.5 magrittr_2.0.3
[5] ggbeeswarm_0.7.2 farver_2.1.1
[7] rmarkdown_2.25 BiocIO_1.12.0
[9] zlibbioc_1.48.0 vctrs_0.6.4
[11] memoise_2.0.1 Rsamtools_2.18.0
[13] DelayedMatrixStats_1.24.0 RCurl_1.98-1.12
[15] htmltools_0.5.6.1 S4Arrays_1.2.0
[17] progress_1.2.2 curl_5.1.0
[19] BiocNeighbors_1.20.0 SparseArray_1.2.0
[21] sass_0.4.7 bslib_0.5.1
[23] cachem_1.0.8 GenomicAlignments_1.38.0
[25] igraph_1.5.1 mime_0.12
[27] lifecycle_1.0.3 pkgconfig_2.0.3
[29] rsvd_1.0.5 R6_2.5.1
[31] fastmap_1.1.1 GenomeInfoDbData_1.2.11
[33] shiny_1.7.5.1 digest_0.6.33
[35] colorspace_2.1-0 dqrng_0.3.1
[37] irlba_2.3.5.1 ExperimentHub_2.10.0
[39] RSQLite_2.3.1 beachmat_2.18.0
[41] labeling_0.4.3 filelock_1.0.2
[43] fansi_1.0.5 httr_1.4.7
[45] abind_1.4-5 compiler_4.3.1
[47] bit64_4.0.5 withr_2.5.1
[49] BiocParallel_1.36.0 viridis_0.6.4
[51] DBI_1.1.3 biomaRt_2.58.0
[53] rappdirs_0.3.3 DelayedArray_0.28.0
[55] bluster_1.12.0 rjson_0.2.21
[57] tools_4.3.1 vipor_0.4.5
[59] beeswarm_0.4.0 interactiveDisplayBase_1.40.0
[61] httpuv_1.6.12 glue_1.6.2
[63] restfulr_0.0.15 promises_1.2.1
[65] grid_4.3.1 Rtsne_0.16
[67] cluster_2.1.4 generics_0.1.3
[69] gtable_0.3.4 hms_1.1.3
[71] metapod_1.10.0 ScaledMatrix_1.10.0
[73] xml2_1.3.5 utf8_1.2.4
[75] XVector_0.42.0 ggrepel_0.9.4
[77] BiocVersion_3.18.0 pillar_1.9.0
[79] stringr_1.5.0 limma_3.58.0
[81] later_1.3.1 dplyr_1.1.3
[83] lattice_0.22-5 rtracklayer_1.62.0
[85] bit_4.0.5 tidyselect_1.2.0
[87] locfit_1.5-9.8 Biostrings_2.70.0
[89] knitr_1.44 gridExtra_2.3
[91] bookdown_0.36 ProtGenerics_1.34.0
[93] edgeR_4.0.0 xfun_0.40
[95] statmod_1.5.0 stringi_1.7.12
[97] lazyeval_0.2.2 yaml_2.3.7
[99] evaluate_0.22 codetools_0.2-19
[101] tibble_3.2.1 BiocManager_1.30.22
[103] graph_1.80.0 cli_3.6.1
[105] xtable_1.8-4 munsell_0.5.0
[107] jquerylib_0.1.4 Rcpp_1.0.11
[109] dir.expiry_1.10.0 png_0.1-8
[111] XML_3.99-0.14 parallel_4.3.1
[113] ellipsis_0.3.2 blob_1.2.4
[115] prettyunits_1.2.0 sparseMatrixStats_1.14.0
[117] bitops_1.0-7 viridisLite_0.4.2
[119] scales_1.2.1 purrr_1.0.2
[121] crayon_1.5.2 rlang_1.1.1
[123] cowplot_1.1.1 KEGGREST_1.42.0 References
Bach, K., S. Pensa, M. Grzelak, J. Hadfield, D. J. Adams, J. C. Marioni, and W. T. Khaled. 2017. “Differentiation dynamics of mammary epithelial cells revealed by single-cell RNA sequencing.” Nat Commun 8 (1): 2128.